pacman::p_load(corrplot, tidyverse, ggstatsplot, ggtern, plotly, seriation, dendextend, heatmaply, GGally, parallelPlot)In-class Exercise 05
Correlogram
wine <- read_csv("data/wine_quality.csv")pairs(wine[,1:11])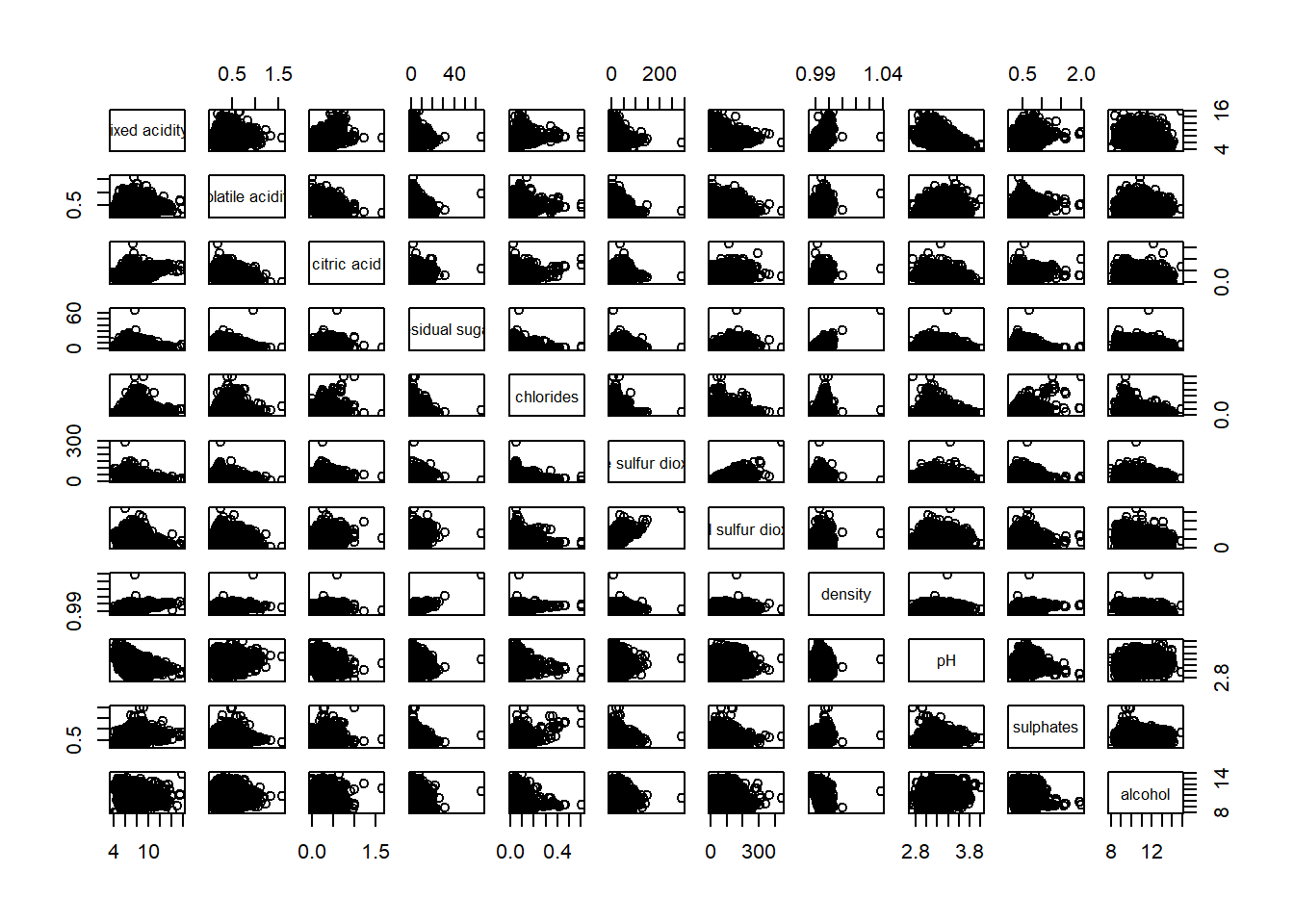
pairs(wine[,2:12])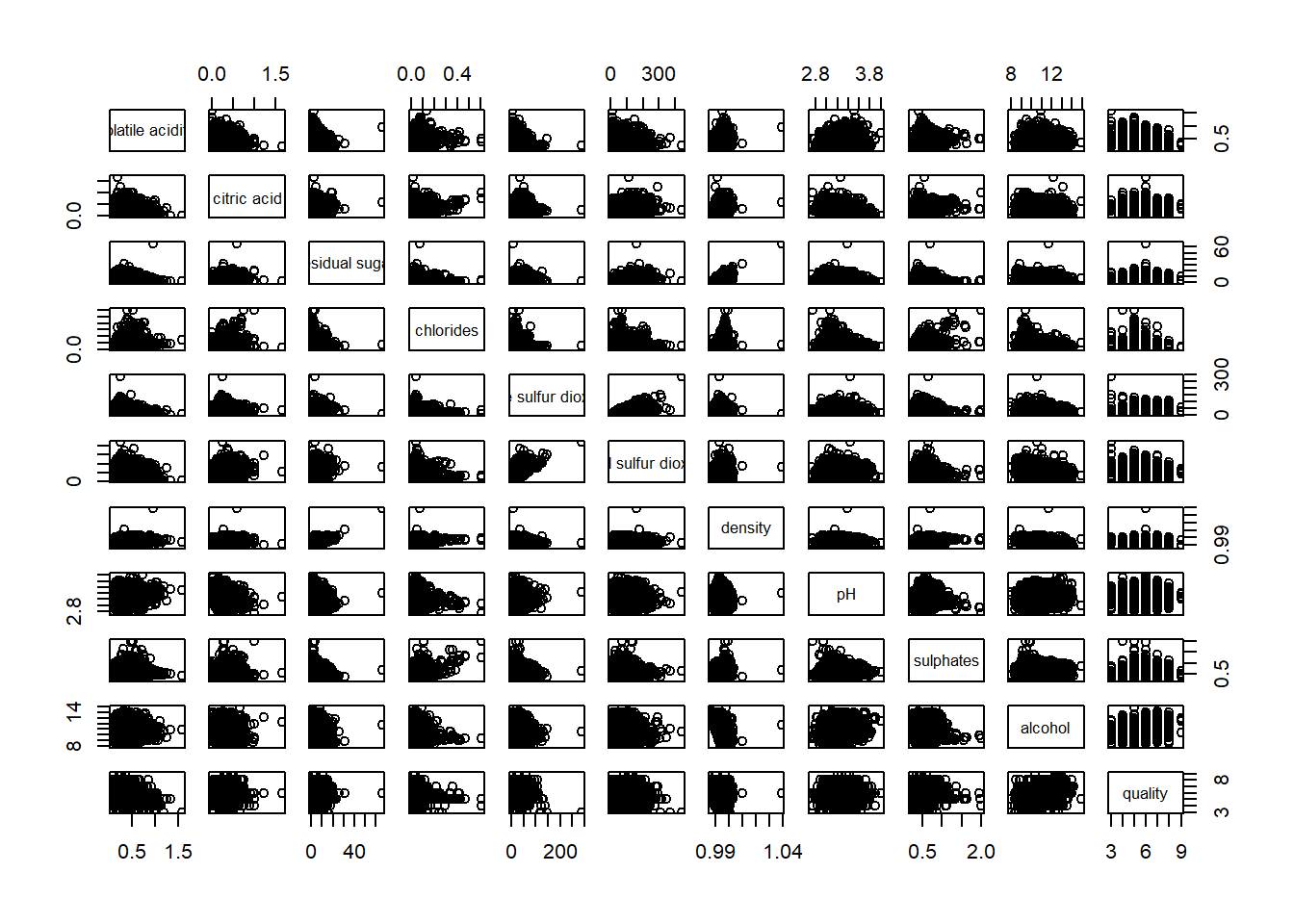
pairs(wine[,2:12], upper.panel = NULL)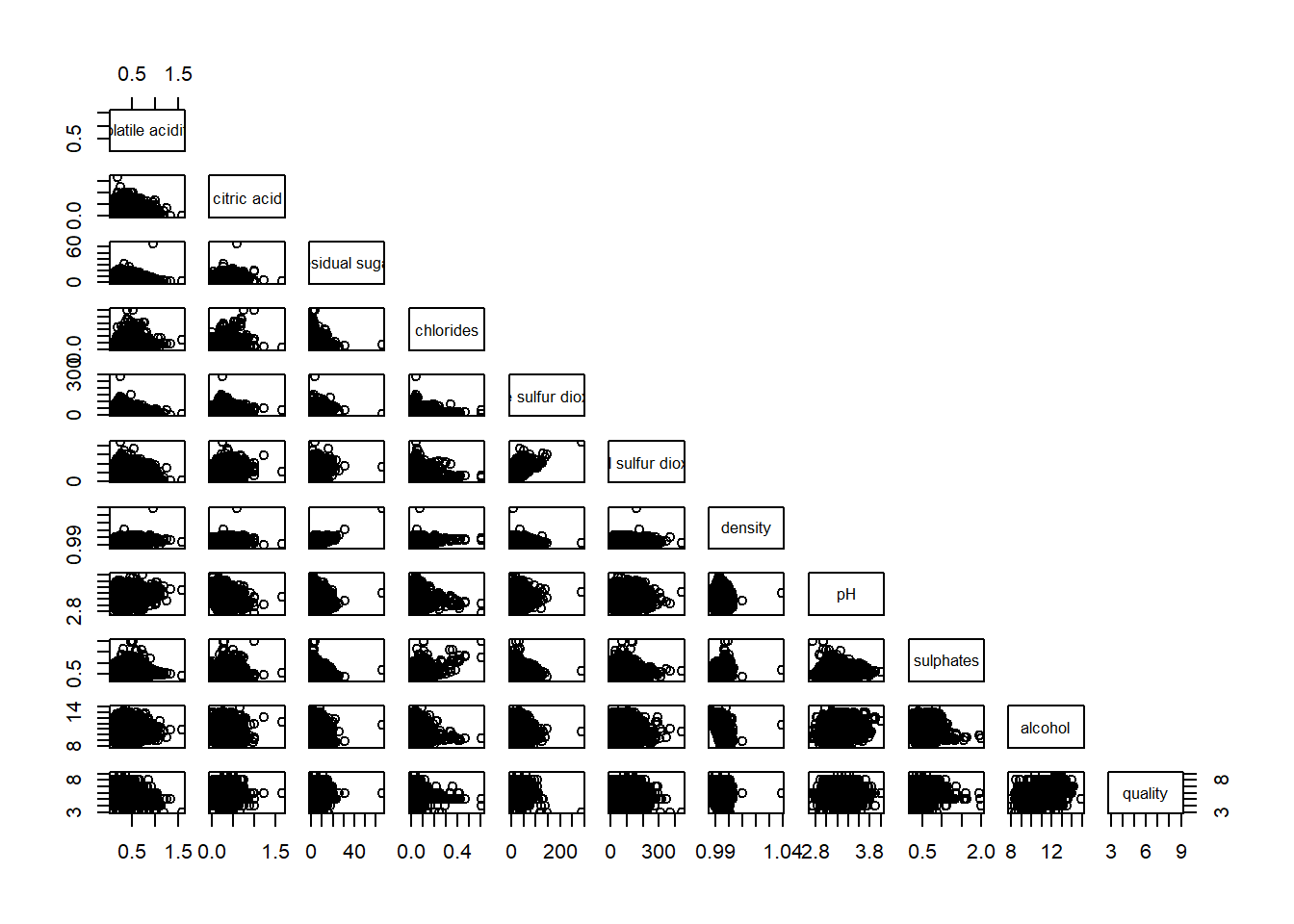
pairs(wine[,2:12], lower.panel = NULL)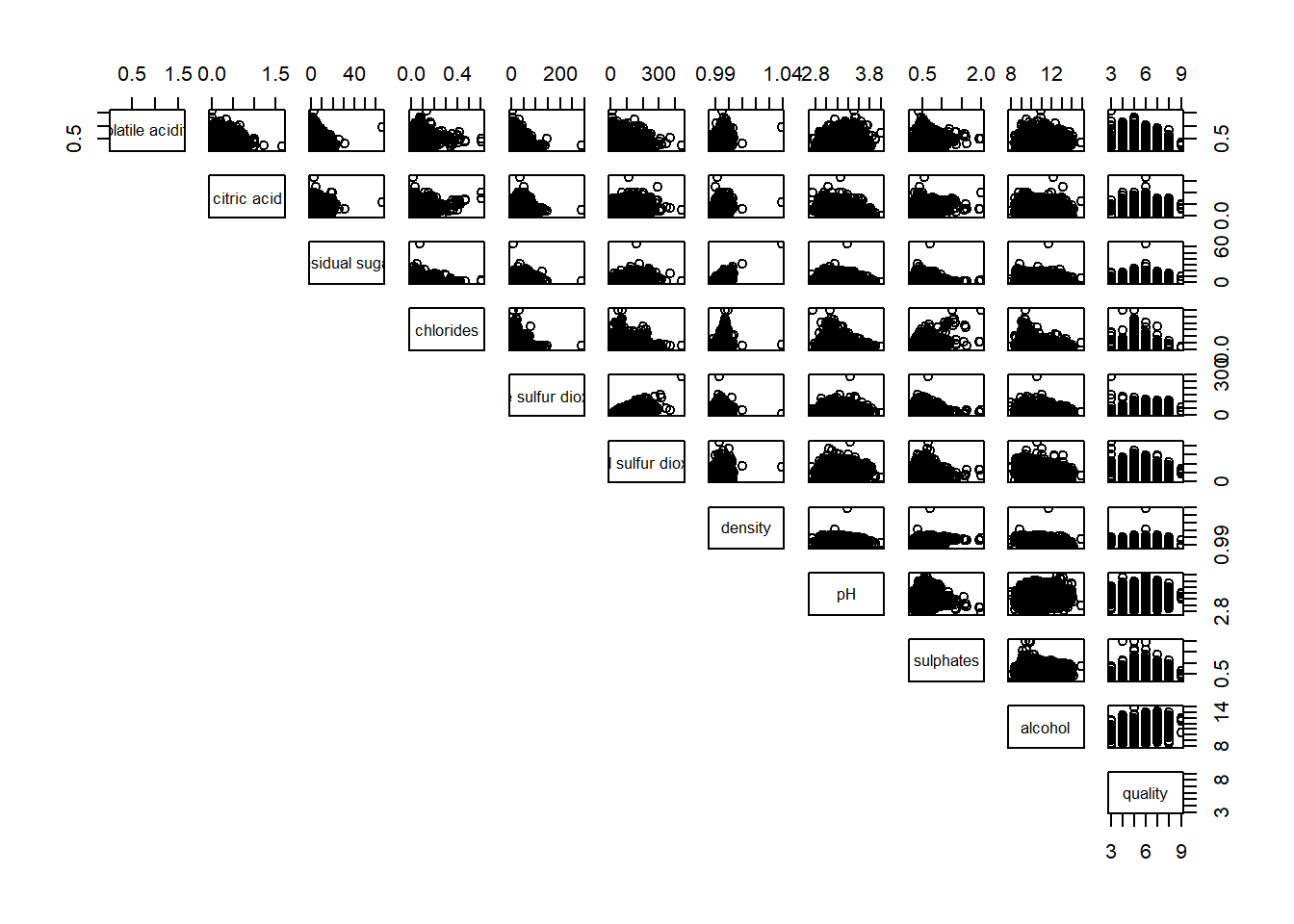
panel.cor <- function(x, y, digits=2, prefix="", cex.cor, ...) {
usr <- par("usr")
on.exit(par(usr))
par(usr = c(0, 1, 0, 1))
r <- abs(cor(x, y, use="complete.obs"))
txt <- format(c(r, 0.123456789), digits=digits)[1]
txt <- paste(prefix, txt, sep="")
if(missing(cex.cor)) cex.cor <- 0.8/strwidth(txt)
text(0.5, 0.5, txt, cex = cex.cor * (1 + r) / 2)
}
pairs(wine[,2:12],
upper.panel = panel.cor)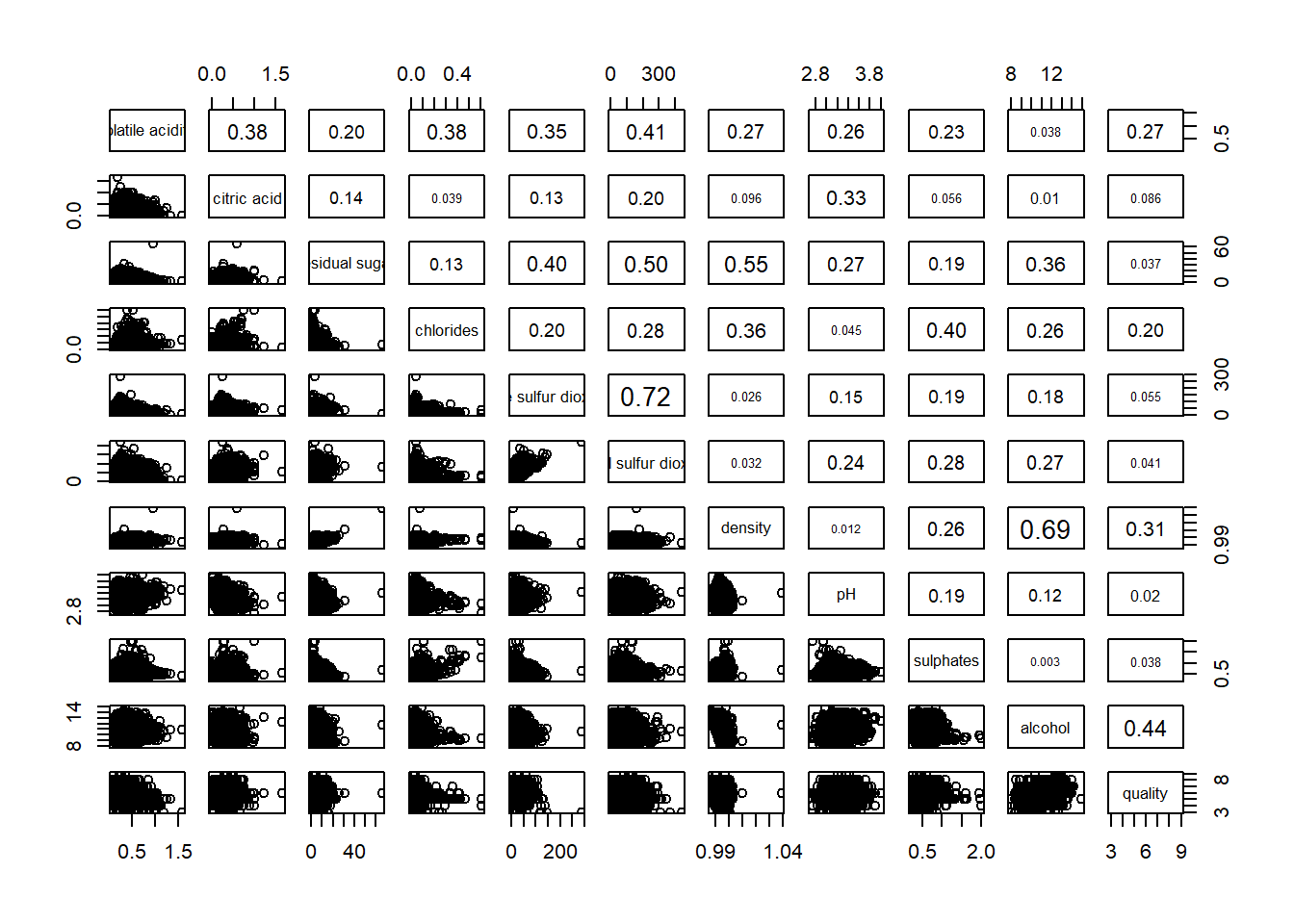
##| fig-width: 7
##| fig-height: 7
#ggstatsplot::ggcorrmat(
# data = wine,
# cor.vars = 1:11,
# ggcorrplot.args = list(outline.color = "black",
# hc.order = TRUE,
# tl.cex = 10,
# lab_size = 3),
# title = "Correlogram for wine dataset",
# subtitle = "Four pairs are no significant at p < 0.05"
#)wine.cor <- cor(wine[, 1:11])corrplot.mixed(wine.cor,
lower = "ellipse",
upper = "number",
tl.pos = "lt",
diag = "l",
tl.col = "black")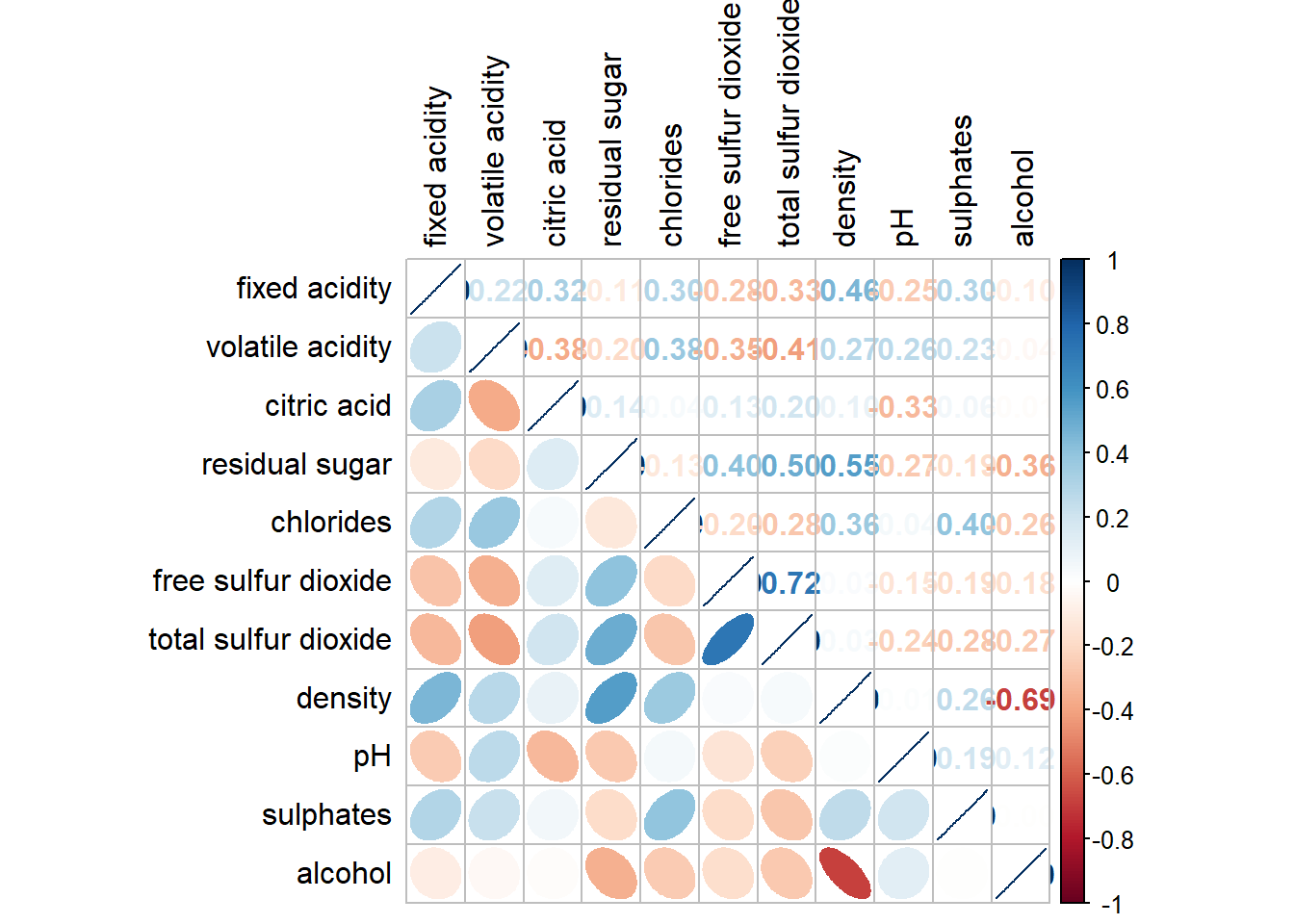
Ternary
pop_data <- read_csv("data/respopagsex2000to2018_tidy.csv") agpop_mutated <- pop_data %>%
mutate(`Year` = as.character(Year))%>%
spread(AG, Population) %>%
mutate(YOUNG = rowSums(.[4:8]))%>%
mutate(ACTIVE = rowSums(.[9:16])) %>%
mutate(OLD = rowSums(.[17:21])) %>%
mutate(TOTAL = rowSums(.[22:24])) %>%
filter(Year == 2018)%>%
filter(TOTAL > 0)ggtern(data=agpop_mutated,aes(x=YOUNG,y=ACTIVE, z=OLD)) +
geom_point()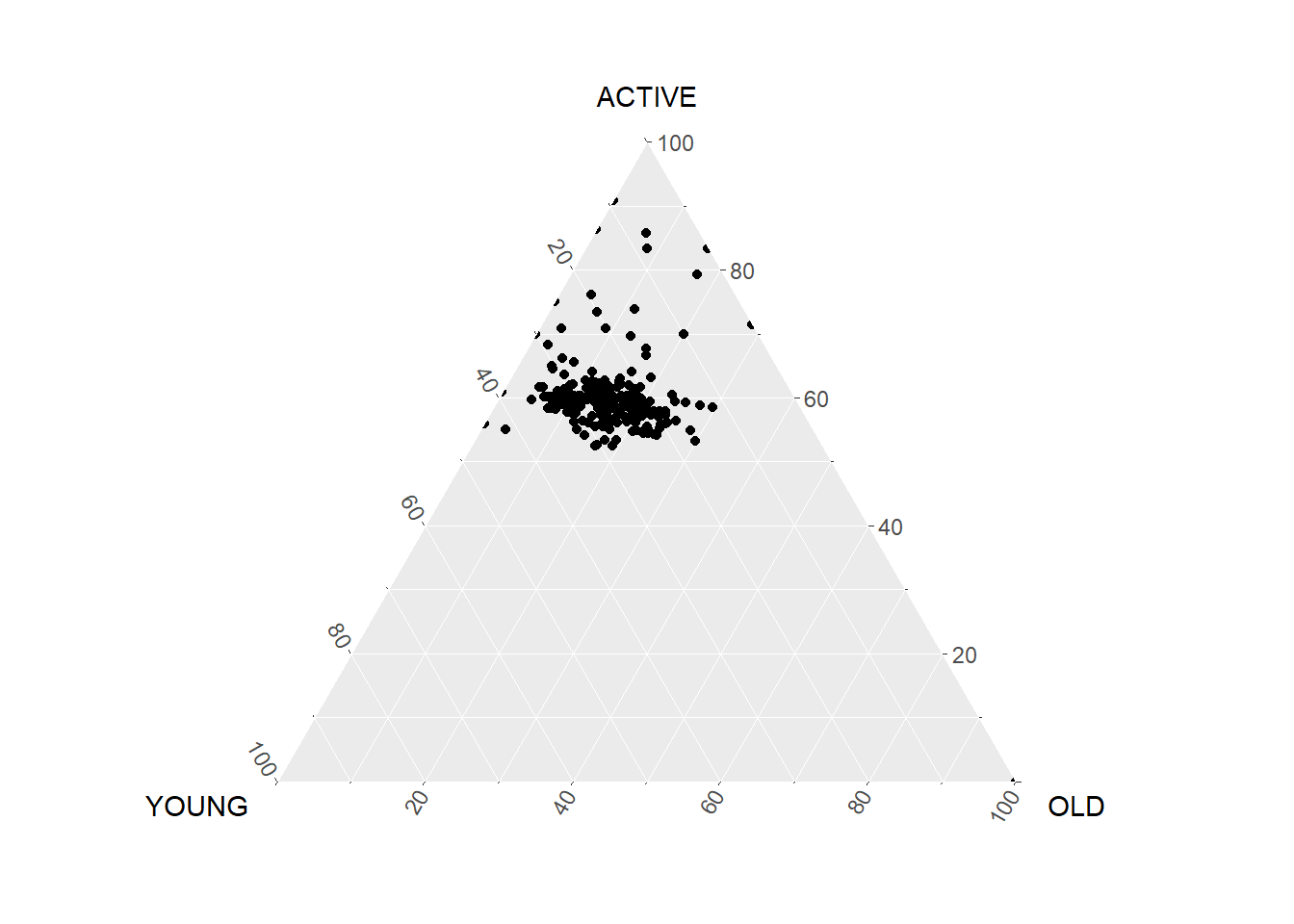
ggtern(data=agpop_mutated, aes(x=YOUNG,y=ACTIVE, z=OLD)) +
geom_point() +
labs(title="Population structure, 2015") +
theme_rgbw()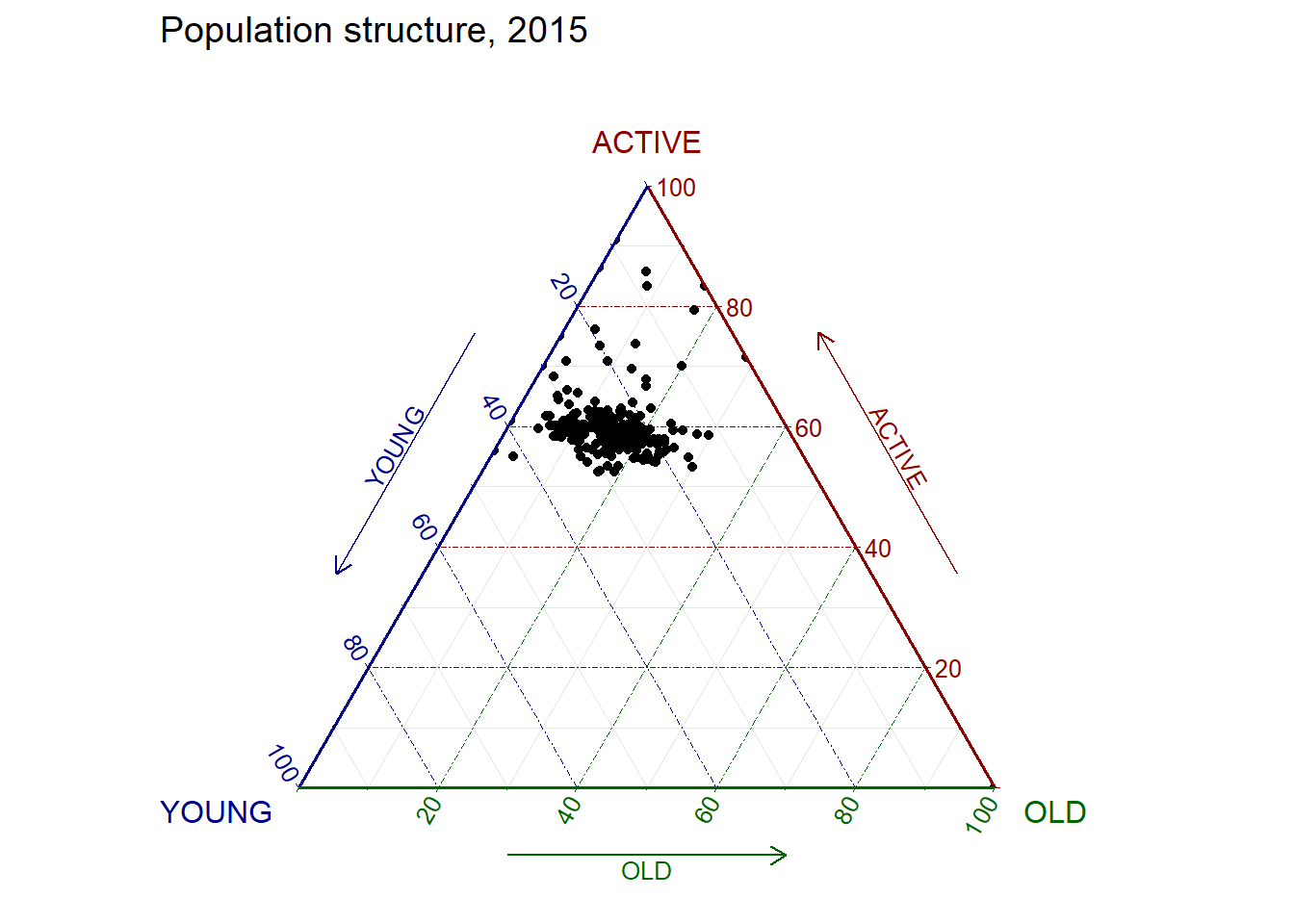
Heatmap
wh <- read_csv("data/WHData-2018.csv")row.names(wh) <- wh$Countrywh1 <- dplyr::select(wh, c(3, 7:12))
wh_matrix <- data.matrix(wh)wh_heatmap <- heatmap(wh_matrix,
Rowv=NA, Colv=NA)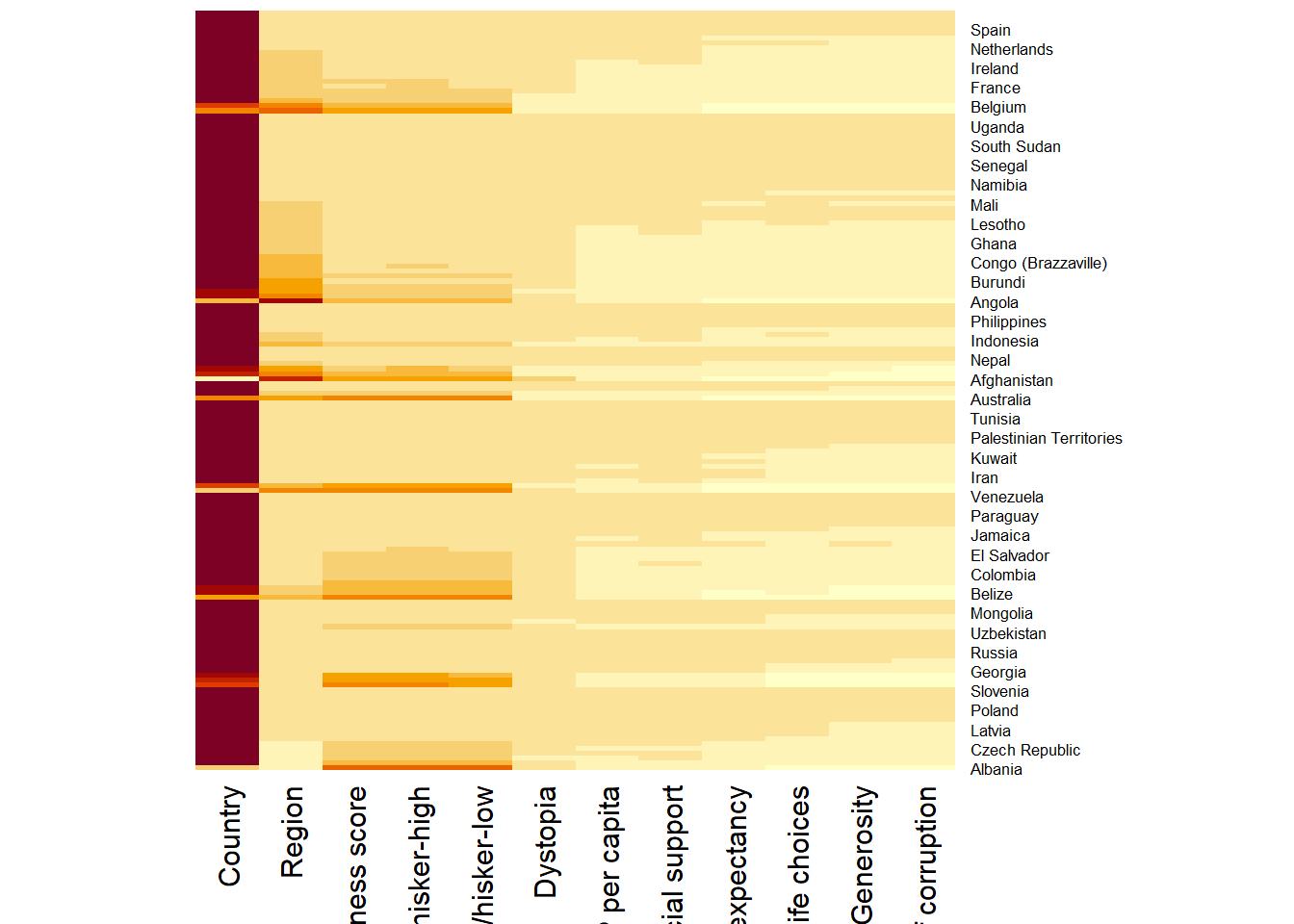
wh_heatmap <- heatmap(wh_matrix)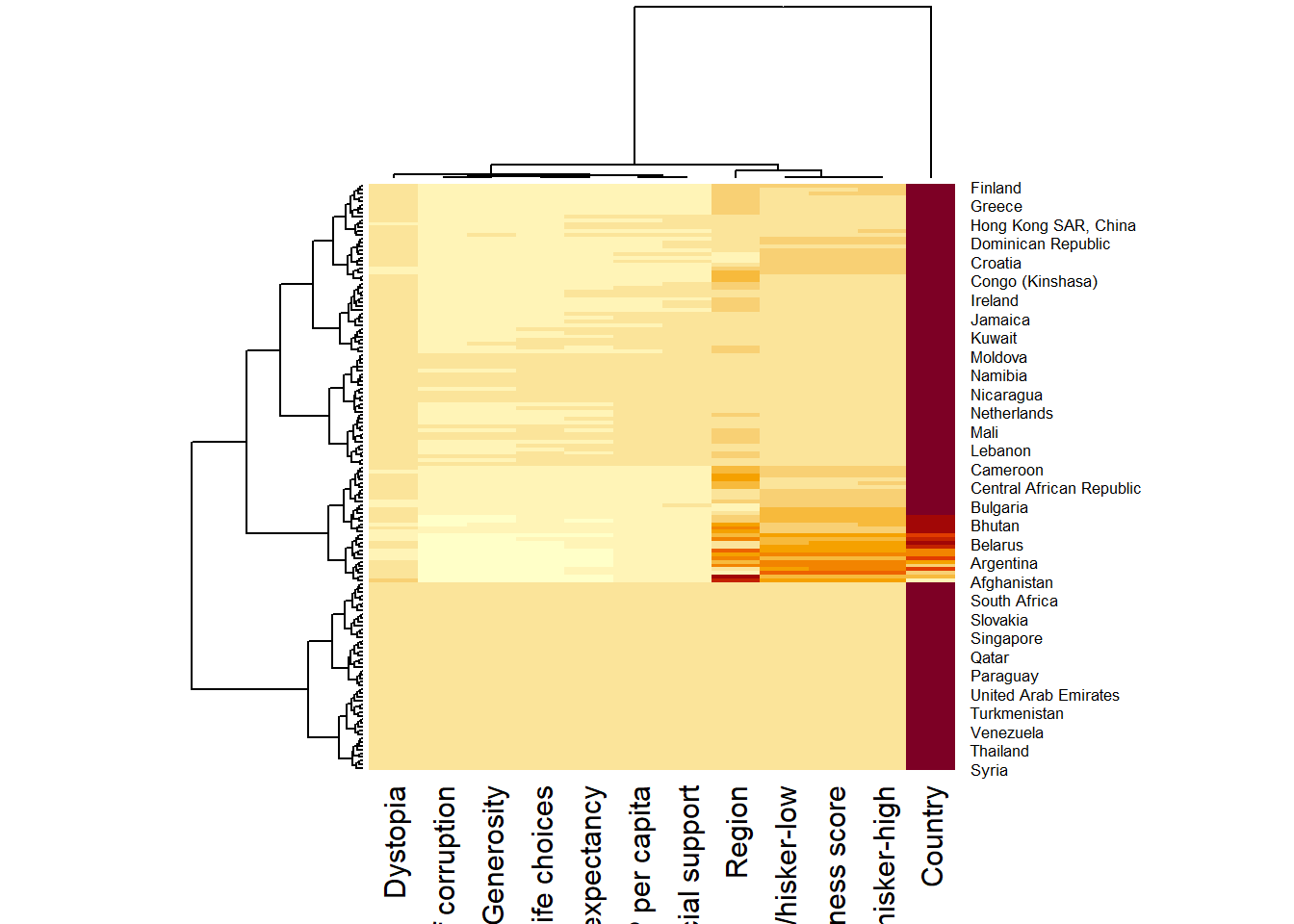
wh_heatmap <- heatmap(wh_matrix,
scale="column",
cexRow = 0.6,
cexCol = 0.8,
margins = c(10, 4))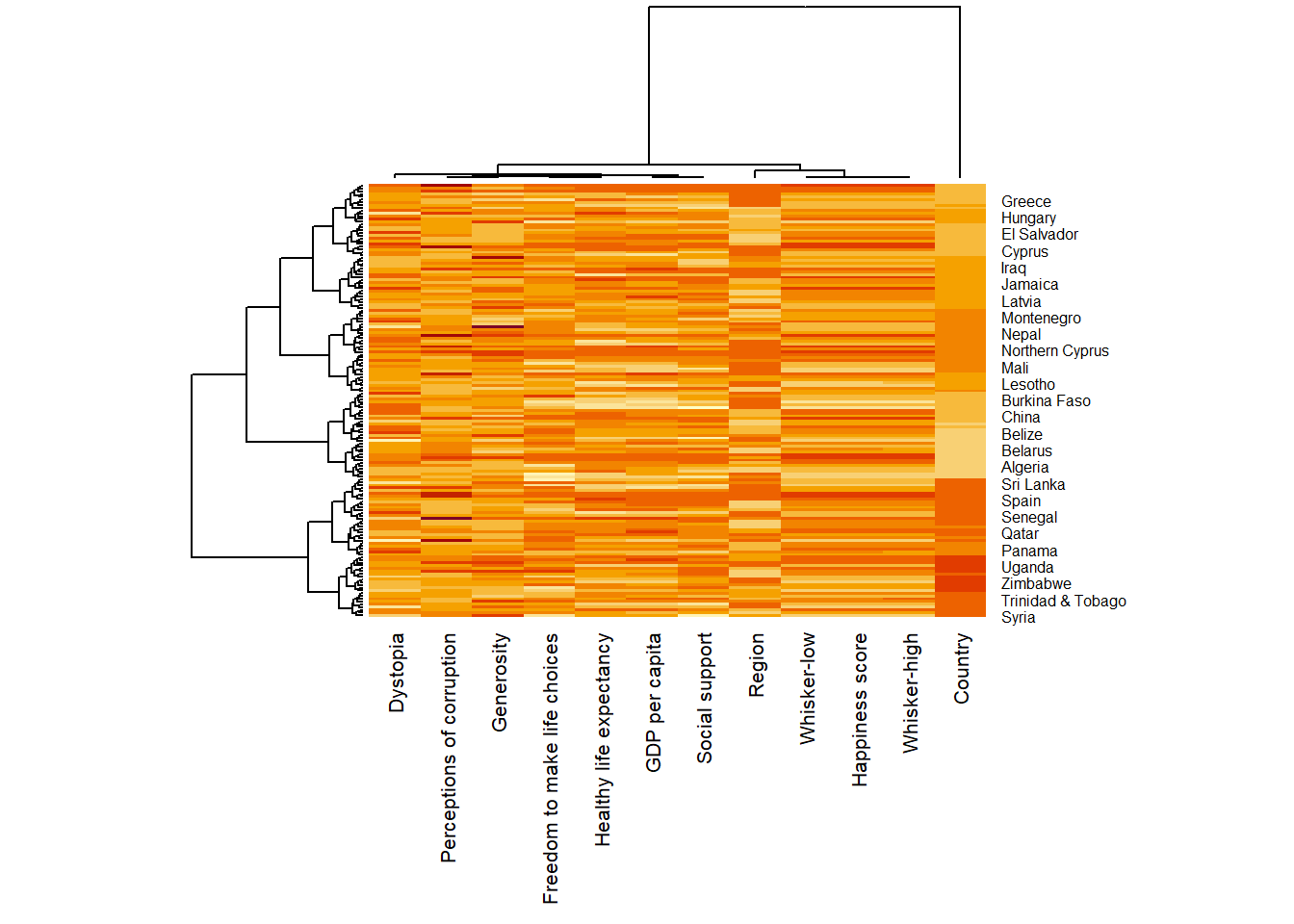
heatmaply(mtcars)heatmaply(wh_matrix[, -c(1, 2, 4, 5)])heatmaply(wh_matrix[, -c(1, 2, 4, 5)],
scale = "column")heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]))heatmaply(percentize(wh_matrix[, -c(1, 2, 4, 5)]))heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
dist_method = "euclidean",
hclust_method = "ward.D")heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
Colv=NA,
seriate = "none",
colors = Blues,
k_row = 5,
margins = c(NA,200,60,NA),
fontsize_row = 4,
fontsize_col = 5,
main="World Happiness Score and Variables by Country, 2018 \nDataTransformation using Normalise Method",
xlab = "World Happiness Indicators",
ylab = "World Countries"
)Parallel Coordinates Plots
wh <- read_csv("data/WHData-2018.csv")ggparcoord(data = wh,
columns = c(7:12))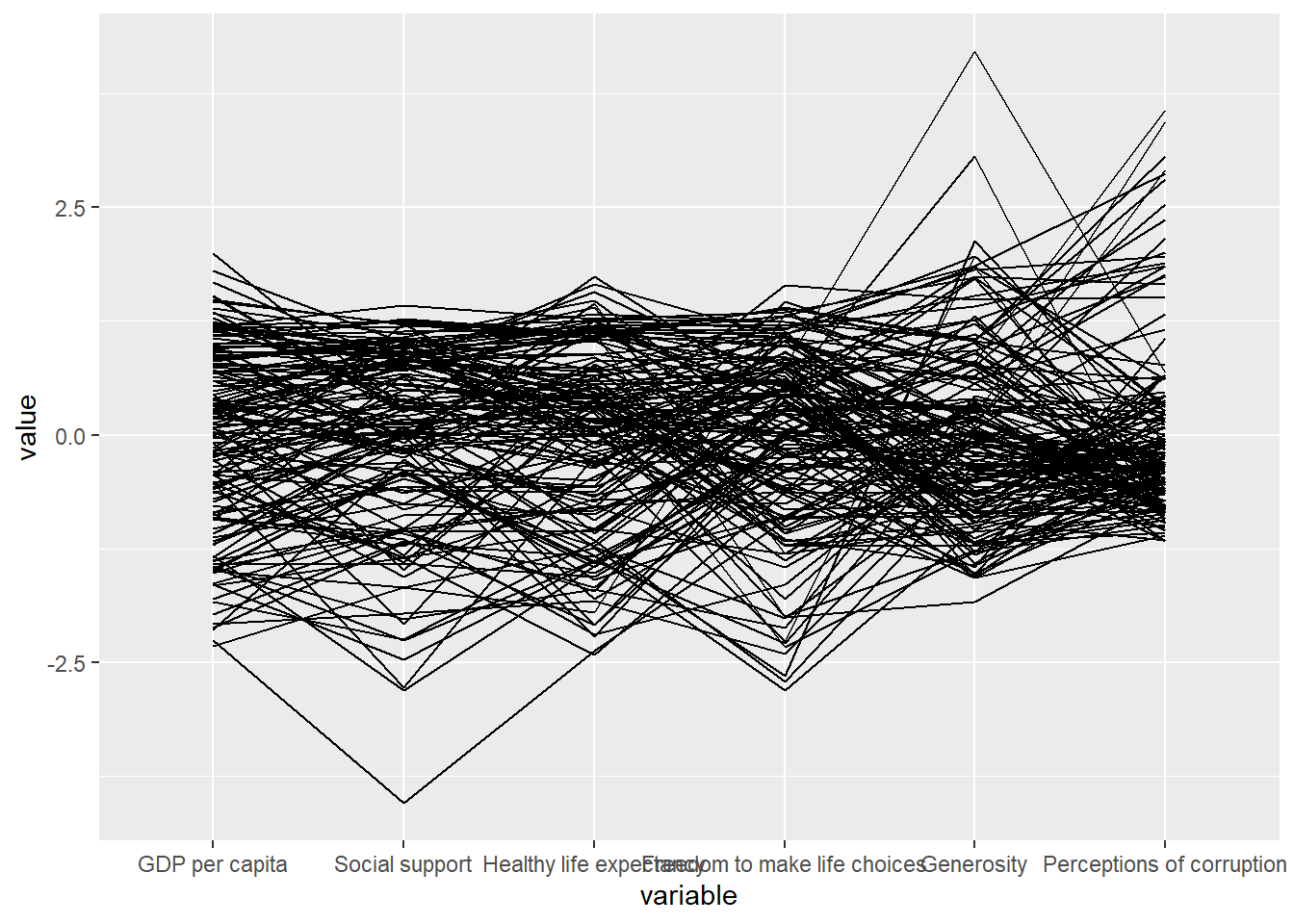
ggparcoord(data = wh,
columns = c(7:12),
groupColumn = 2,
scale = "uniminmax",
alphaLines = 0.2,
boxplot = TRUE,
title = "Parallel Coordinates Plot of World Happines Variables")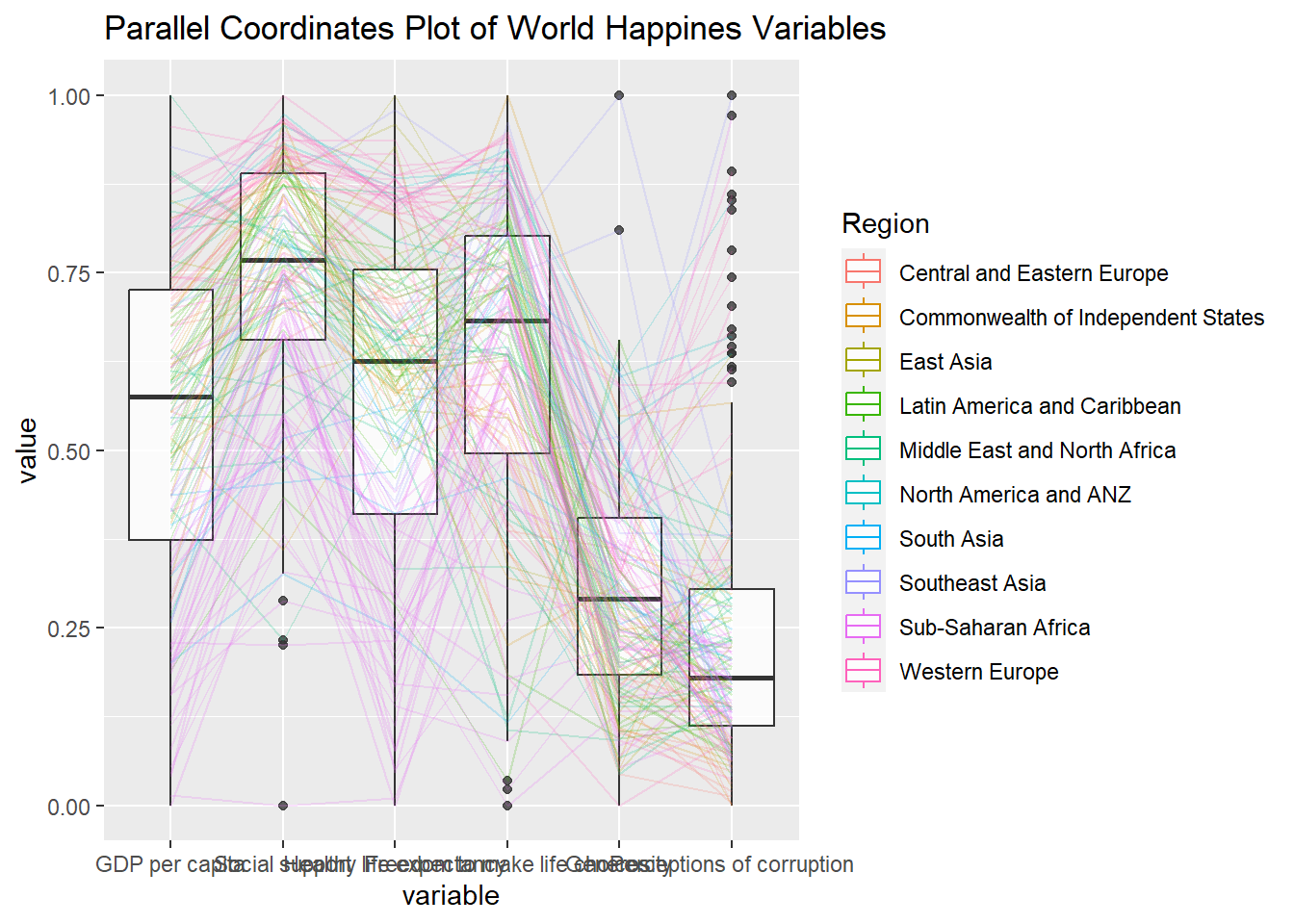
ggparcoord(data = wh,
columns = c(7:12),
groupColumn = 2,
scale = "uniminmax",
alphaLines = 0.2,
boxplot = TRUE,
title = "Multiple Parallel Coordinates Plots of World Happines Variables by Region") +
facet_wrap(~ Region)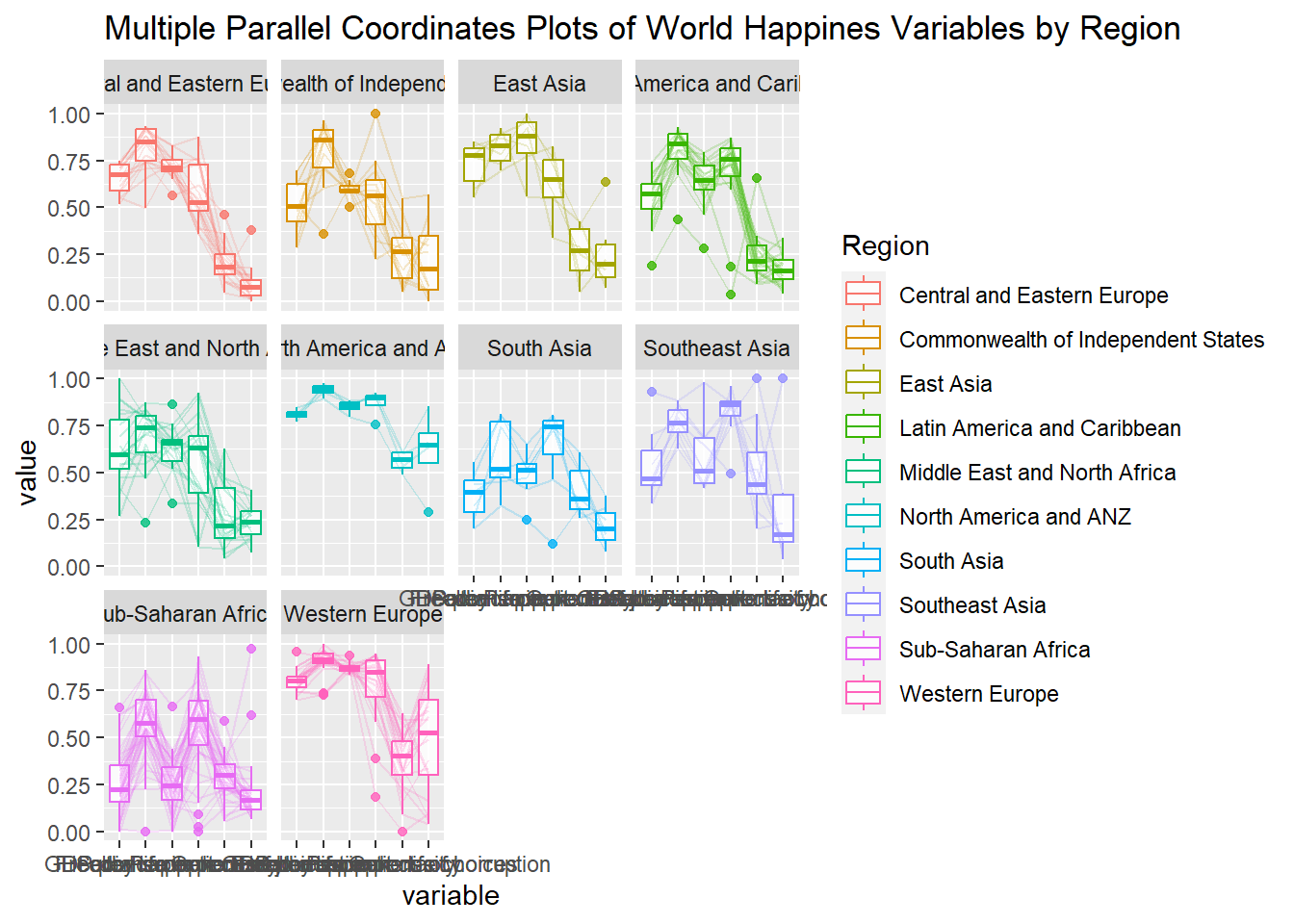
histoVisibility <- rep(TRUE, ncol(wh))
parallelPlot(wh,
rotateTitle = TRUE,
histoVisibility = histoVisibility)